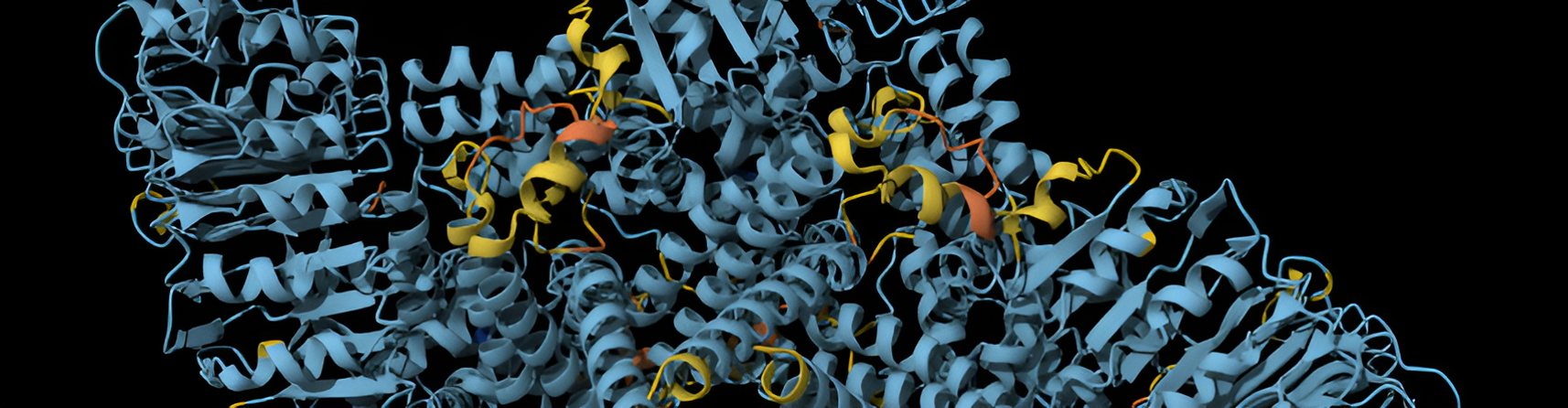
NLR Evolution in A. thaliana and its Relatives
The evolution of NLR resistance genes in Arabidopsis thaliana and its relatives reflects adaptation to microbial threats, with these resistance genes key to detecting pathogen effectors and triggering immunity. We investigate the ecological, genomic, and functional drivers of resistance gene evolution by studying NLR-effector interactions across hosts and populations.
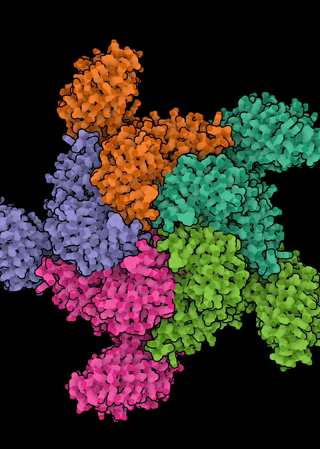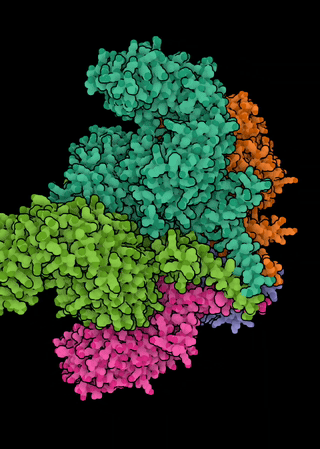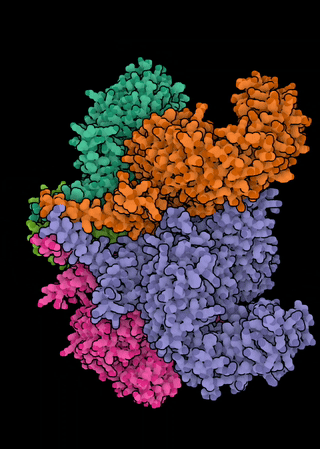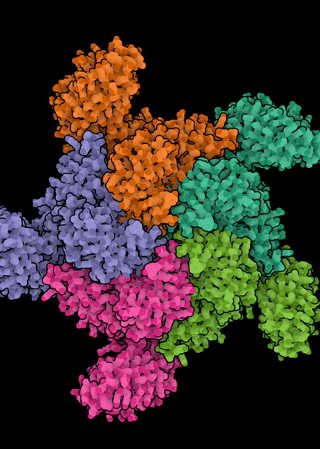
The NLR genes in A. thaliana exhibit a range of evolutionary histories, from ancient balanced polymorphisms through neutrality to selective sweeps. Since many of the pathogens that attack A. thaliana are generalists, and both the microbiota and pathobiota of Arabidopsis are shared with related and co-occurring hosts, we are interested in the extent to which molecular evolutionary dynamics of resistance play out at the community level. In particular, we ask: can we see evidence of congruent molecular evolution at orthologous NLR resistance genes across A. thaliana, Cardamine hirsuta, and Draba verna? We are further interested in the extent of functional conservation of orthologous NLRs among these hosts—especially in whether the repertoires of recognized effectors diverge and whether they are similarly integrated within the broader NLR immune signaling network.This effort focuses on natural populations in which these three hosts co-occur across a metapopulation in the Midi-Pyrénées region of southern France. This project allows a dual focus on both the importance of ecological context on the evolution of resistance genes and on the evolution of the immune system itself.
- Maerkle, H and J Bergelson. Evolutionary implications of host interactions with a generalist pathogen. In review.
- Weiner, B, Maerkle, H, Laderman, E, Demirjian, C and J Bergelson. 2025. A physical model links structure and function in the plant immune system. PNAS, in press.
- Karasov, TL, Kniskern, JM, Gao, L, DeYoung, BJ, Ding, J, Ullrich, D, Lastra, R, Nallu, S, Innes, RW, Barrett, LG, Hudson, RR and J Bergelson. 2014. The long-term maintenance of a resistance polymorphism through diffuse interactions. Nature 512 (7515): 436–440.